The FAIRVASC Challenge and Opportunity
Rigorous clinical research relies upon analysis of a sufficiently large number of observations (patients, samples, experimental measurements, etc.) to allow reliable statistical inference. The low prevalence of rare diseases means that patient numbers are small in any given country. The resulting small cohort sizes represent a major barrier to such research. While national and local registries are emerging for many rare diseases, federating them so that they can be used as a single pool of data has three key challenges – finding the registry with the data needed (registry discovery), overcoming variations in the data held, how it is structured and the terminologies used (data normalisation) and legal, ethical and regulatory constraints on accessing and using the data (data governance). The FAIRVASC workplan consists of eight work-packages (WPs).
AAV: a model autoimmune rare disease, with applicability in other rare disease arenas: The challenge of fragmented and heterogeneous registries applies across many rare diseases in Europe and beyond. In FAIRVASC, we focus on the rare disease ANCA-associated vasculitis (AAV, ORPHA:156152 ) as a demonstrator of how to use RDF and semantic technologies to normalise and access registry data across borders.
The development of the FAIRVASC infrastructure is an iterative process, which involves the interaction of three teams established as part of the project. These are: the Harmonisation Implementation Team (HIT), the FAIRVASC Implementation Team (FIT) and the Query Implementation Team (QIT).
The HIT team is made up of ANCA-associated vasculitis (AAV) experts from each registry who give feedback on proposed harmonised clinical terms from the various FAIRVASC registries. Once agreed, these harmonised terms are formally integrated into the FAIRVASC ontology by the Work Package 3 (WP3) team.
The FIT team is made up of IT professionals who have been trained in the task of uplifting the registry data into the linked data format RDF, based on the developed ontology. The FIT and HIT teams meet every two weeks (alternately). The HIT team reviews and agrees upon recent ontology development, then the following week this new aspect of the ontology is presented to the FIT team for technical implementation at their registry site. A small team of three individuals representing Work Packages 1, 3 and 5 coordinates this process and designs the initial harmonisation structure to present to the HIT team.
The QIT team works on the co-design of the federate queries which will be run over the registry data. These queries will ultimately by run on real patient data at the local registry site and will return irrevocably anonymised, aggregated data on vasculitis patients across Europe.
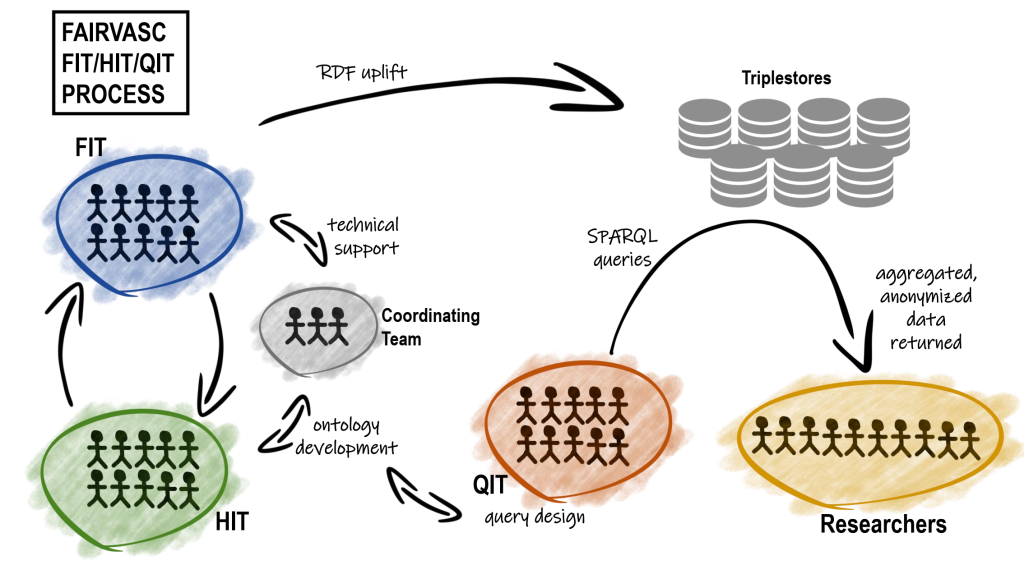
The workflow of FAIRVASC project and interactions between teams:
The original instantiation of the QIT team developed the first set of experimental questions or queries of key importance to FAIRVASC.
Based on these questions, the HIT team analyse the registries and co-design the ontology. The work of the HIT is than followed by FIT team, which develop R2RML mappings for all registries. R2RML is a semantic web technology which allows tabular data to be transformed, or ‘uplifted’ into linked data.
When the original QIT queries are satisfied, the QIT team will once again begin to co-design an extended set of SPARQL queries, to explore new aspects of vasculitis which are of interest to researchers. SPARQL is a ‘query language’ for linked data – it allows questions to be asked of the FAIRVASC infrastructure, once the data has been uplifted.
Due to the aggregation of patient data from various European registries planned in the next stages of FAIRVASC, a Data Quality Team was created. Its task is to develop criteria and methods for assessing the quality of the collected data.
To disseminate the project progress and results there are dedicated work packages (WPs) realized by Communication and Dissemination team to communicate the FAIRVASC concept with target audiences including academia, industry, rare disease patients and patient advocacy organisations (PAOs), policymakers and the public.
